 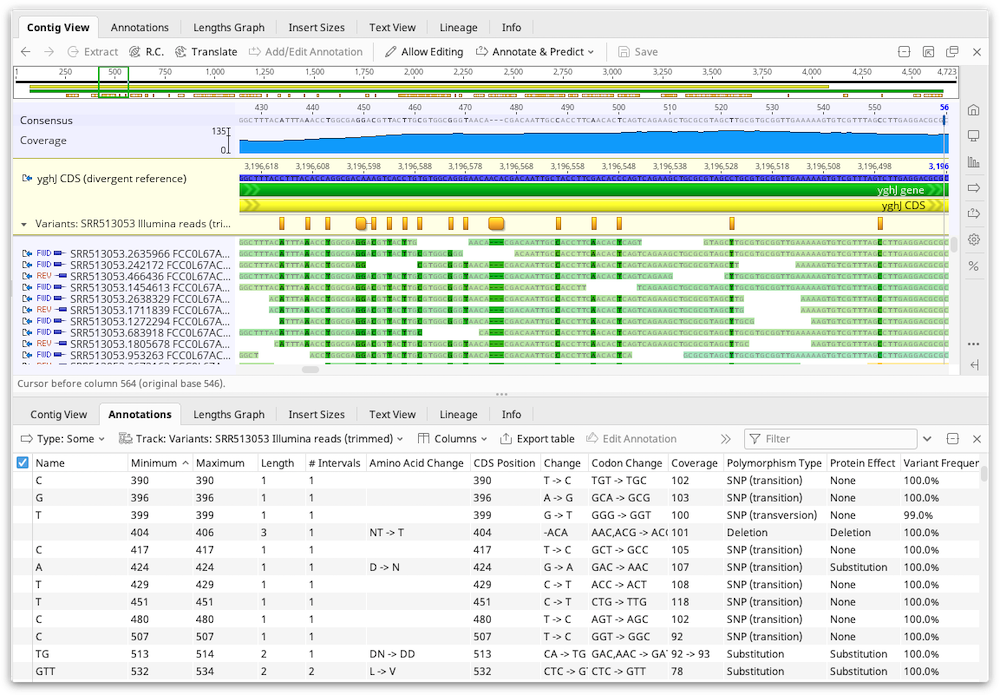
We now support specifying custom liabilities and assets for specific points on IgG-like sequences, for example IMGT position 118F on the light chain. Positional Liabilities based on antibody numbering schemes You can also open your documents in a new tab quickly by selecting this button on the preview page: When selecting a document, the preview window allows you to inspect a summary of the documents, as well as quickly preview the sequences and tables.Ĭhoosing Open Full Document immediately opens the document in a wide-view, as shown below: We are very excited to show off our new user interface for accessing documents in Geneious Biologics. Please let us know if you use Microsoft Azure/Google authentication to login to Geneious Biologics, and are also interested in a Geneious Biologics/Geneious Prime integration. ** Our Geneious Prime connector plugin is currently available for non-SSO customers only. A new analysis option ideal for larger datasets.Ability to flexibly add clusters and graphs to existing analysis results ( replaces the more limited "Recluster" pipeline).Streamlined Reference Database creation, including a reference sequence auto-formatter, and increased support for different database types (Germline Genes, target/scaffold variable regions, and other features).We'd love to hear about what formats you are using in your work. If you are interested in this feature, or support for other types or multivalent/ multispecific formats, please get in touch. Our first phase will add support for dual nanobody sequences (two VHH sequences on one read). The old version is no longer in support, please see our information about how to update.** New speedier integration with Geneious Prime.Clustering options now available in Single Clone Antibody Analysis.Adding labels to a selection of sequences.Positional Liabilities based on antibody numbering schemes.Open documents in full view quickly and easily.Preview documents to see the history and info.An improved user interface experience allowing you to:.The number of raw trees saved will therefore be equal to the number of samples.Over the past few months we have been working on some exciting new features for Geneious Biologics. If this is turned on then all of the trees created during resampling will be save in the resulting tree document. 
The percentage of topologies in the original trees which must be represented by the summarizing topologies. This is used to decide which monophyletic clades to include in the consensus tree, after comparing all the trees in the original set. Produce trees which summarise the topologies resulting from resampling. Choose this to create a consensus tree from the resampled data. 
The number of alignments and trees to generate while resampling. Either bootstrapping or jackknifing can be performed when resampling columns of the sequence alignment. Choose which sample to use as an outgroup, or leave it as “no outgroup” to build an unrooted tree. There are two methods under this option – Neighbor joining and UPGMA. If you are building a tree from amino acid sequences you only have the option of “Jukes Cantor” distance correction. If you are building a tree from DNA sequences you have the choices “Jukes Cantor”, “HKY” and “Tamura Nei”. This lets the user choose the kind of substitution model used to estimate branch lengths. Excludes sites containing Masked annotations from the analysis without permanently removing them from your alignment. For more information on these options see sections 12.3 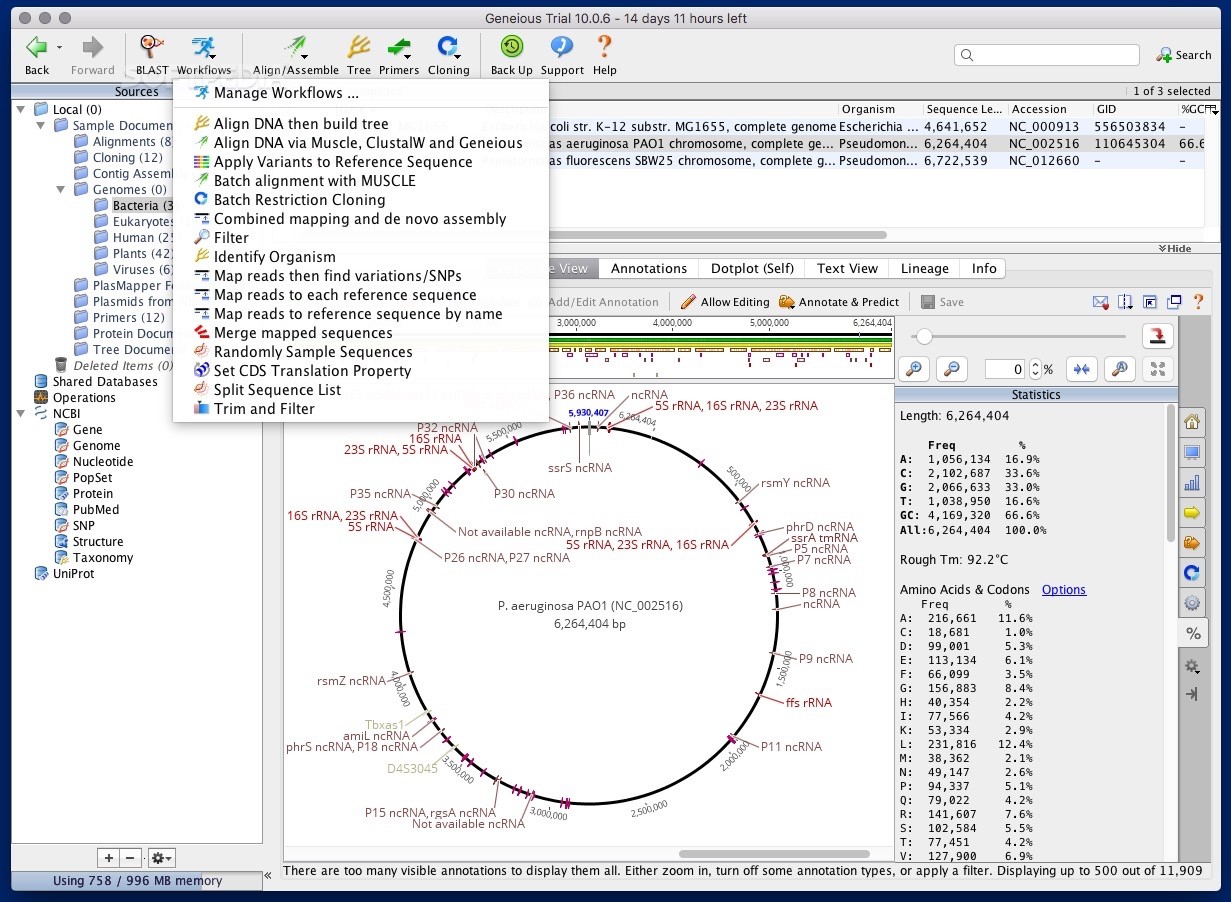
The following options are available in the tree-building dialog for the Geneious tree builder. Figure 12.2: Tree building options in Geneious Prime 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
